Collyer's second claim is that "[n]atural selection, the supposed basis of evolution,
can only select from existing characteristics and does not produce new genetic
material." Both statements are wrong. Natural selection is not the only mechanism of evolutionary change. Genetic drift is another mechanism of evolutionary change, and is of particular importance at the molecular level where much of the sequence difference among species is likely due to drift.
His appalling grasp of evolutionary biology is demonstrated by his claim that natural selection 'does not produce new genetic material.' Of course it doesn't! It is not the mechanism by which new genetic material appears. Point mutation, genetic duplication, chromosomal duplication, insertion of mobile genetic elements, genome duplication, lateral gene transfer and endosymbiosis are the mechanisms by which new genetic material appears in the genome. Natural selection, as the mechanism of adaptive change, then acts on these genetic changes.
Natural selection acts on new genetic diversity, which is produced by a number of mechanisms:
- Point mutation: this is the most common form of mutation, and occurs when the nucleotide at a site in the genome is replaced by a different nucleotide. Some mutations are silent; others can result in a faulty gene product, while others can be beneficial.
- Insertion and deletion: these mutations occur when one or more nucleotides are inserted into, or deleted from a gene. Amino acids are encoded by three consecutive nucleotides, so if these insertions are not multiples of three, the result will be a frameshift mutation that will result in a truncated gene.
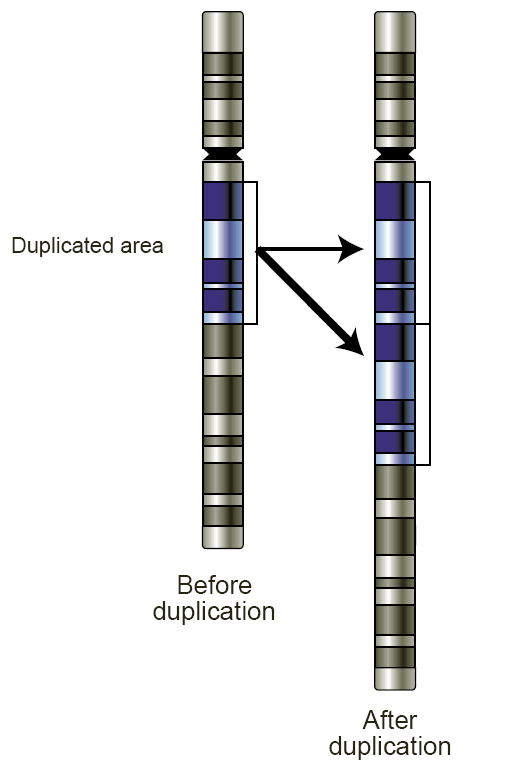
- Gene duplication: occasionally, genes (or even whole chromosomes) can be copied, resulting in more than one copy. Mutations in the extra copy can result in a decayed, non-functional gene (pseudogene) or it can result in a gene with a different function, which can allow new functionality to evolve. [1]
- Whole genome duplication: genomic analysis of many species shows that duplication of the whole genome has occurred in the remote past.
- Insertion of transposons: these are a class of mobile genetic elements that copy and paste, or cut and paste themselves from one part of the genome to another part. Retrotransposons copy themselves via an RNA intermediate, while DNA transposons achieve this transposition without the need for an RNA intermediate. Transposition can cause disease, but has also been linked to the evolution of new functionality.
- Integration of proviral sequences into host genomes: retroviruses, unlike DNA viruses encode their genetic material in RNA. They are able to insert a DNA copy of themselves into the host genome. If they insert themselves into the germline, they can be passed onto the next generation. Depending on where they insert themselves, they can alter the expression of nearby genes. Elements of the proviral sequence can be co-opted by the host genome for completely different uses.
A fascinating example of how endogenous retroviral elements can be co-opted by evolution for completely new purposes comes from the IRG gene family, which plays a part in the immune system. In apes and monkeys, this gene family had been reduced to a single gene which was then crippled by the insertion of a retrotransposon. The authors of the study note how the chance integration of a retroviral element, which acted as a promoter, along with two further mutation events restored the IRGM gene in the common ancestor of apes and humans. [2]
- Horizontal gene transfer: the transfer of genetic elements from one species to another is quite common in bacteria, but has been noted in metazoans. The transferring of antibiotic resistance between different bacterial species is one notable example of horizontal gene transfer.
- Nature Geoscience 6, 688–690 doi:10.1038/ngeo1939
Some of these changes are deleterious, while some are
beneficial. Many other are neutral and may convey an advantage to the
population should environmental conditions change later. Contrary to the
special creationist distortion of modern evolutionary biology, there are a
number of ways in which beneficial mutations occur on which natural selection
can work.
Furthermore, natural selection acting on genetic novelty is not the only mechanism of evolutionary change. Genetic drift occurs due to random genetic fluctuations, and unlike natural selection is not an adaptive mechanism of evolutionary change. Douglas Futuyma notes that:
Furthermore, natural selection acting on genetic novelty is not the only mechanism of evolutionary change. Genetic drift occurs due to random genetic fluctuations, and unlike natural selection is not an adaptive mechanism of evolutionary change. Douglas Futuyma notes that:
Genetic drift and natural selection are the two most important causes of allele substitution - that is, of evolutionary change - in populations. Genetic drift occurs in all natural populations because, unlike ideal populations at Hardy-Weinberg equilibrium, natural populations are finite in size. Random fluctuations in allele frequencies can result in the replacement of old alleles by new ones, resulting.in nonadaptive evolution. That is, while natural selection results in adaptation, genetic drift does not-so this process is not responsible for those anatomical, physiological, and behavioral feahu·es of organisms that equip them for reproduction and survival. Genetic drift nevertheless has many important consequences, especially at the molecular genetic level: it appears to account for much of the difference in DNA sequences among species. [3]
An excellent example of how these mechanisms create new genetic information is the evolution ofan antifreeze gene found in Antarctic notothenioid fish from an ancient trypsinogen gene:
The study authors provide the most likely mechanism by which an ancestral gene was converted into an antifreeze protein gene:
The 5′ end (E1, I1, and small segment of E2) and the 3′ end (I5 3′ splice site and E6) of trypsinogen gene were recruited and linked, and the remainder of the gene deleted (dashed lines and boxes). The Thr-Ala-Ala coding element was duplicated, presumably via slippage at the repetitive (gt)n sequence during replication. The recruited E1 provided the 5′ UTR and signal peptide sequences for the new AFGP gene. The deletion, linking, and amplification events led to a 1-nt frameshift resulting in a termination codon (tga) at the start of the recruited trypsinogen E6 and converting it into the 3′ flanking sequence of the AFGP gene. The spacer sequence (bars filled with zigzagged lines) and additional I1 sequence might be existing sequence in the trypsinogen progenitor gene or acquired through recombinatory events. The Thr-Ala-Ala coding duplicants plus a spacer became amplified de novo to form the new AFGP polyprotein coding region. [4]
Conclusion
Collyer's assertion is wrong on two accounts. He ignored the fact that natural selection is not the only mechanism of evolutionary change. Genetic drift plays an important role at the molecular level. The main blunder was his assertion that 'natural selection does not produce new genetic material.' It acts on mutation, gene duplication, horizontal gene transfer, and most definitely results in the addition of new genetic material.
References
1. An excellent overview of how gene duplication results in the creation of genetic information can be found here: Gene Duplication versus ID Panda's Thumb May 23 2004
2. Bekpen C, Marques-Bonet T, Alkan C, Antonacci F, Leogrande MB, et al. (2009) Death and Resurrection of the Human IRGM Gene. PLoS Genet 5(3):
e1000403.
doi:10.1371/journal.pgen.1000403
3. Futuyma D "Evolution" (2005, Sinauer Associates ) p 226
4.Chen L et al “Evolution of anti-freeze glycoprotein from a trypsinogen gene in Antarctic notothenoid fish” Proceedings of the National Academy of Science (1997) 94:3811-16